Deep learning to achieve clinically applicable segmentation of head and neck anatomy for radiotherapy
12 Sep 2018We propose a model for segmenting the organs at risk in 3D Computed Tomography (CT) scans for the planning of radiotherapy treatment. Improvements in such a segmentation could result in improved and faster treatment.
arXiv
Open-sourced augmentation
Open-sourced surface distance metrics
Stanislav Nikolov, Sam Blackwell, Ruheena Mendes, Jeffrey De Fauw, Clemens Meyer, Cían Hughes, Harry Askham, Bernardino Romera-Paredes, Alan Karthikesalingam, Carlton Chu, Dawn Carnell, Cheng Boon, Derek D’Souza, Syed Ali Moinuddin, Kevin Sullivan, DeepMind Radiographer Consortium, Hugh Montgomery, Geraint Rees, Ricky Sharma, Mustafa Suleyman, Trevor Back, Joseph R. Ledsam, Olaf Ronneberger
Abstract
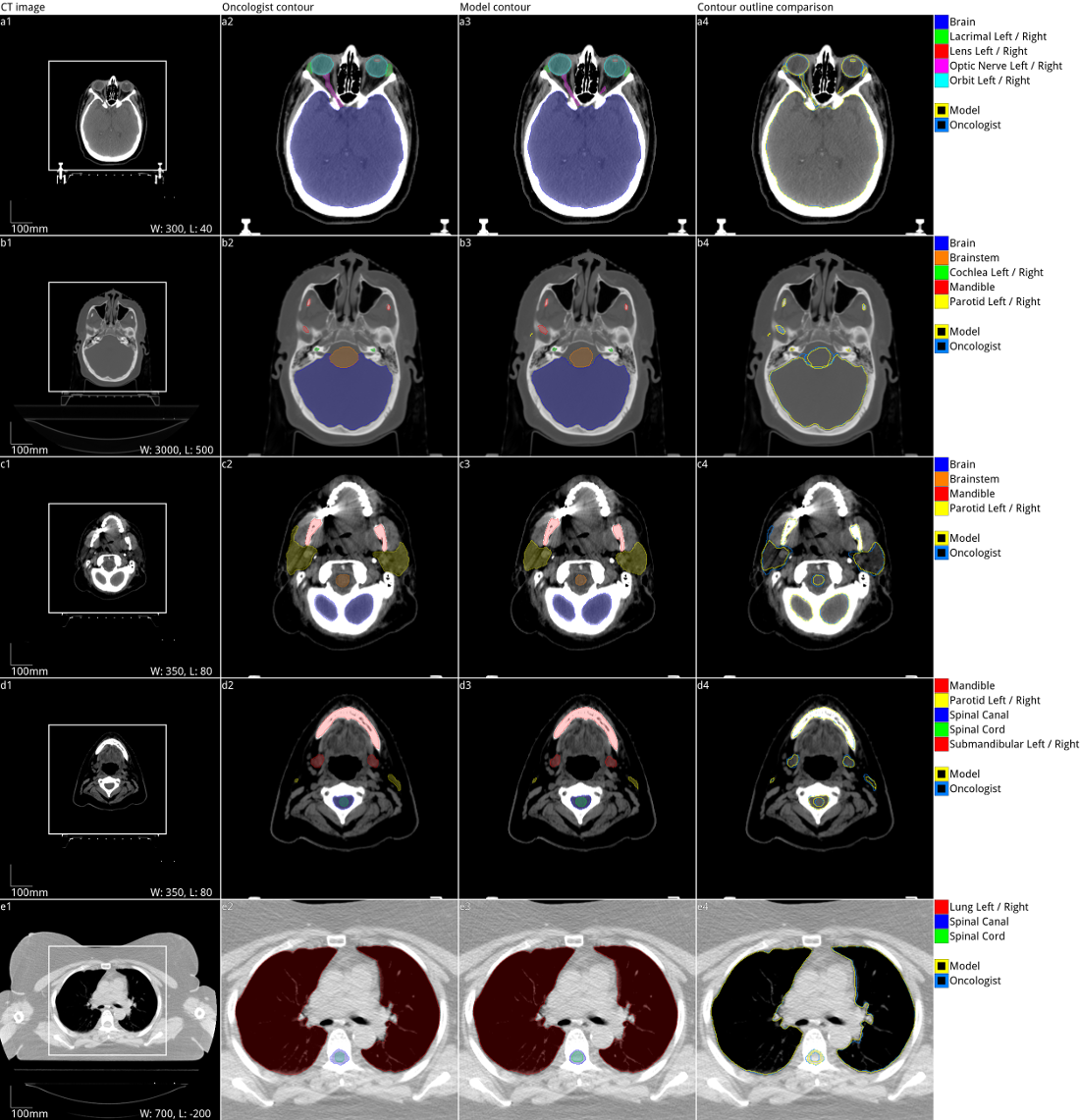
Over half a million individuals are diagnosed with head and neck cancer each year worldwide. Radiotherapy is an important curative treatment for this disease, but it requires manually intensive delineation of radiosensitive organs at risk (OARs). This planning process can delay treatment commencement. While auto-segmentation algorithms offer a potentially time-saving solution, the challenges in defining, quantifying and achieving expert performance remain. Adopting a deep learning approach, we demonstrate a 3D U-Net architecture that achieves performance similar to experts in delineating a wide range of head and neck OARs. The model was trained on a dataset of 663 deidentified computed tomography (CT) scans acquired in routine clinical practice and segmented according to consensus OAR definitions. We demonstrate its generalisability through application to an independent test set of 24 CT scans available from The Cancer Imaging Archive collected at multiple international sites previously unseen to the model, each segmented by two independent experts and consisting of 21 OARs commonly segmented in clinical practice. With appropriate validation studies and regulatory approvals, this system could improve the effectiveness of radiotherapy pathways.